Notice that an object "protein" was added to the right-hand task bar. create protein, chain A and chain B and chain C). If a protein has more than one chain, you should include all of them (e.g. Create an object for your protein by typing into the PyMOL command line (either in the Tcl/Tk GUI, or in the Viewer), the following PyMOL command: create protein, chain A.Visually locate the bound ligand in the structure. Type color yellow, resn G39 to color the bound ligand in a distinct color. Color is very useful for identifying ligands.It is rather hard to identify the ligand in this representation. Slowly rotate the molecule on the screen and try to see where the bound ligand is.From Plugin, select PDB Loader Service, and enter 2HT8.Follow these instructions to quickly locate the substrate binding pocket in influenza neuraminidase. This is a fairly trivial task if a structure of the protein-ligand complex is available from the PDB. This is the most powerful way to interact with the program, but requires that you are familiar with the logic and syntax of PyMOL commands.Īs most drugs bind at the surface of the protein, it is a common task to visualize the protein surface in order to locate the binding pocket. For example, you can create dashed lines (a good way to show hydrogen bonds) between oxygens and nearby amide hydrogens in the alpha-helix by using a command "dist (helix and name o), (helix and (name h and neighbor name n)), 2.5". The command line allows you to type in commands that can accomplish almost any of PyMOL's functions.For example, you can select two atoms (first by CTRL-SHIFT/left-clicking, second by CTRL-SHIFT/right-clicking ) and create a covalent bond by pressing CTRL-T. Furthermore, you can select and perform specific tasks via mouse and keyboard shortcuts. Your three-button mouse allows you to rotate, move, and zoom in/out by holding down the left, middle, or right button, respectively. The molecular display area allows to one view and interactively manipulate molecules.
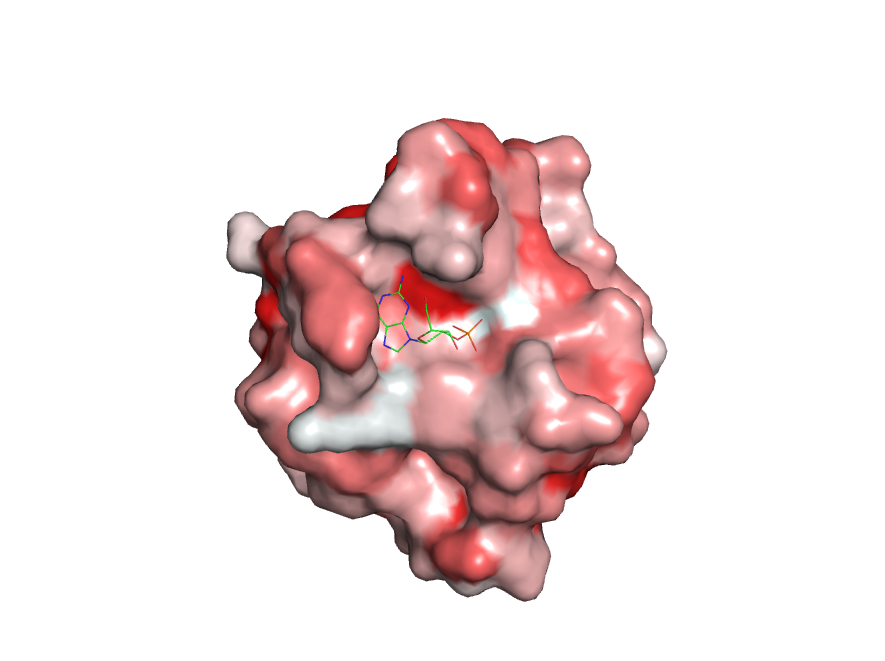
For example, you can show your protein as a grey surface and the inhibitor as green sticks if you have previously separated them into two different objects.
#Pymol tutorial wiki how to
#Pymol tutorial wiki install
PyMOL is available for many computer platforms, and students in the drug design course may install a precompiled educational versions (available from the course instructor) to their personal computers. In more difficult cases you can ask help from PyMOL users via PyMOL forum. Examples of PyMOL images, with commands used to generate these views, are available in the Biomolecular Images and Movies for Teaching page.
#Pymol tutorial wiki manual
The PyMOL commands are explained in the PyMOL manual and in PyMOL Wiki. But the menu-driven user interface is not particularly intuitive and it takes a little time to learn the program. As a result, a powerful functionality is available via PyMOL commands that one types into the PyMOL command line. Some of the plugins, such as ABPS for the calculation of electrostatic potentials in proteins, CAVER for identifying access routes to protein cavities, and LigAlign for atom-to-atom mapping of ligands in different structures are useful in structure-based drug design.ĭuring the PyMOL development, emphasis has been on functionality and not on ease of use. The strengths of PyMOL arise from the emphasis on high-quality graphics, the use of the powerful programming language ( Python), and the extendability with Plugins.

It can also perform many other valuable tasks (such as editing PDB files) to assist you in your research. Warren DeLano, PyMOL is a molecular graphics system with an embedded Python interpreter designed for real-time visualization and rapid generation of high-quality molecular graphics images and animations. According to the program's author, late Dr. We will be using PyMOL for visualizing macromolecules. Tutorial: Molecular Visualization of Protein-Drug InteractionsĬomputer-Aided Drug Design Tutorials: Visualization of Protein Surfaces with PyMOL PyMOL for Molecular Visualization

